Software and databases
Unless stated otherwise, the files shown below are © Miguel Á. Carreira-Perpiñán. They are free for non-commercial use. I make them available in the hope that they will be useful, but without any warranty.
 Github repository
Github repository
Some of the software produced by my students and I is in https://github.com/UCMerced-ML.
For implementations of my own algorithms, see my publications page.
Machine learning algorithms and models
- Algorithms based on the method of auxiliary coordinates (MAC):
- Computation of affinity or similarity functions:
- Dimensionality reduction algorithms:
- Elastic embedding (EE) (reference: Carreira-Perpiñán, ICML 2010). We have three codes:
- Dimensionality reduction by unsupervised regression (DRUR) (reference: Carreira-Perpiñán and Lu, CVPR 2008, 2010).
- lapeig.m: Laplacian eigenmaps (reference: Belkin and Niyogi, Neural Comp. 2003).
- Locally Linear Landmarks (LLL): fast, approximate Laplacian eigenmaps for large-scale manifold learning (reference: Vladymyrov and Carreira-Perpiñán, ECML/PKDD 2013).
- Variational Nyström (VN): fast, approximate Laplacian eigenmaps for large-scale manifold learning (reference: Vladymyrov and Carreira-Perpiñán, ICML 2016).
- Stochastic neighbor embedding (SNE) and t-SNE (references: Hinton and Roweis, NIPS 2002; van der Maaten and Hinton, JMLR 2008).
For small datasets (thousands of points), this code is the fastest one available for SNE and t-SNE as far as I know. It uses the spectral direction (reference: Vladymyrov and Carreira-Perpiñán, ICML 2012). For large datasets, use the fast multipole implementation above.
- Clustering algorithms:
- Semi-supervised learning algorithms:
- Probabilistic models:
- Gaussian mixtures:
- Gaussian mixture Matlab tools contains a set of Matlab functions to perform some common operations with Gaussian mixtures (GMs): computing the density, gradient and Hessian at given points; computing the moments (mean, covariance); finding all the modes (with a fixed-point, mean-shift iteration); sampling from the GM; learning the GM parameters with an EM algorithm; learning a Gaussian classifier; reconstructing missing values in a dataset using a GM; and finding the parameters of a conditional or marginal GM, e.g. for p(x3|x2,x4) from a GM with pdf p(x1,x2,x3,x4,x5). In particular, this allows finding the modes, gradient, etc. of conditional or marginal GMs, with isotropic, diagonal or full covariance matrices.
Here is an application of these tools to reconstructing missing data in a database of articulatory speech (Wisconsin X-ray Microbeam Database): code (reference: Qin and Carreira-Perpiñán, Interspeech 2010).
- Mode-finding in Gaussian mixtures (including the calculation of error bars, gradient and Hessian). See my page on Gaussian mixtures. This code assumes isotropic covariance matrices, for more general covariance matrices and many other useful functions, see the Gaussian mixture Matlab tools in this page.
- Generalised elastic nets (GEN): these extend the original elastic net of Durbin and Willshaw to arbitrary differential operators. This generalised elastic nets web page contains figures and animations for travelling salesman problems (TSP) and visual cortical map simulations of V1, all created with the Generalised Elastic Net Matlab Toolbox (reference: Carreira-Perpiñán and Goodhill, 2003).
- Other functions:
- sqdist.m: matrix of squared Euclidean distances between two datasets (needed by many of the functions below).
- nnsqdist.m: k nearest neighbours (indices and distances) of a dataset.
- knn.m: k-nearest neighbour classifier.
- lagdist.m: lagged distances of a vector time series.
- roc.m: ROC curve for a binary classifier.
- confumat.m: confusion matrix for a K-class classifier.
- recall1.m: recall@R measure in information retrieval: the average rate of queries for which the 1-nearest neighbour is ranked in the top R positions.
- kmeans.m: k-means algorithm, with various choices for the initialisation (including kmeans++).
- kmeans1.m: a generalisation of the k-means algorithm to use weighted centroids and a (partially) fixed codebook.
- procrustes.m: Procrustes alignment of two datasets. Useful to compare the results of different dimensionality reduction algorithms.
- rrr.m and rrrtrain.m: reduced-rank regression (linear regression where the coefficient matrix is constrained to be low-rank).
- gaussaff.m: Gaussian affinity matrix of a dataset based on various types of graphs: full, k-nearest-neighbour (symmetric, nonsymmetric, mutual), epsilon-ball. Returns a sparse matrix if possible.
- seglsq.m: segmented least squares problem (using dynamic programming), i.e., fit several lines to a 2D dataset.
- econncomp.m: connected components of a point set using the Euclidean distance.
- conncomp.m: connected components of a graph.
- sample_dirichlet.m: sampling from a Dirichlet distribution.
- sw.m: Swendsen-Wang Monte Carlo sampling.
- varimax.m: varimax rotation.
- imgsqd.m and imgsqd2.m: transform an image file (greyscale or colour) into a data set with one feature vector per pixel and computes the sparse matrix of squared Euclidean distances between pixel feature vectors (requires rgb2lab.m, rgb2luv.m, rgb2xyz.m). The difference between imgsqd.m and imgsqd2.m is in how boundary pixels are treated: in imgsqd.m their square neighbourhood is clipped, so they have fewer neighbours than interior pixels, while in imgsqd2.m the neighbourhood is shifted inside, so each pixel has the same number of neighbours.
- rgb2lab.m, rgb2luv.m, rgb2xyz.m: color space transformations.
- map50.m: create a 50% saturation colormap, with unobtrusive colours as in political maps, useful to plot the regions created by clustering or classification algorithms with 2D datasets.
Numerical optimisation
The following functions are part of my course materials for EECS260 Optimization:
- numgradhess.m: evaluates numerically (with a finite-difference approximation) the gradient and Hessian of a function at a list of points, and optionally compares these gradients and Hessians with user-provided ones (useful to confirm the latter are correct). See the example function Fquad.m.
- AC.m: compare pairs of arrays numerically using the infinity-norm (useful to verify that two different computations are essentially the same numerically).
- convseq.m: estimates numerically the order of convergence given a sequence of vectors.
- fcontours.m: contour plot of a 2D function with equality and inequality constraints.
- plotseq.m: plots a sequence of points in 2D (useful in combination with fcontours.m).
- Osteepdesc.m: steepest descent minimisation.
- linesearch.m: backtracking line search.
- Example functions: Fquad.m (quadratic), Frosenbrock.m (Rosenbrock function in n dimensions), Fsines.m (sines in 2D).
Some more functions:
- proxbqp.m: solves a collection of proximal bound-constrained quadratic programs each of the form min(x'.A.x - 2b'.x + μ.|x-v|2) s.t. l ≤ x ≤ u. It uses an efficient ADMM algorithm described here.
- SimplexProj.m: computes the projection of a vector onto the simplex (the standard, regular or probability simplex), fully vectorised and efficient (O(n.logn) where n is the dimension of the vector). The algorithm is briefly described here.
(Articulatory) speech processing
- Articulatory databases: the following tools allow you to read data files from the MOCHA and XRMB databases into Matlab, plot animations of the vocal tract (pellet locations and outlines of the tongue and palate) and speech (waveform, spectrogram, energy, pitch, and phoneme labels if available), to save the visualisation as a movie file, to produce scatterplots and temporal plots of the articulator traces, and various other things:
Both of them require you to install our speech analysis functions. This code was mostly written by my PhD student Chao Qin. We also have a Java interface for this code written by Jimmy Yih and Mark Crompton.
- Electropalatography: tools for imaging EPG frames.
 Databases
Databases
 implementations
implementations
Weka is a suite of machine learning software for data mining tasks written in Java. We have implemented the following algorithms in Weka:
- MACPcv: a LaTeX2e class file to typeset a personalised curriculum vitae.
- MACPremark: a LaTeX2e package to add remarks to a draft LaTeX file, such as I need to complete this section and add a bibliographic reference or Extend this proof to the complex numbers.
- MACPpreprint: a LaTeX2e package to add remarks to a preprint LaTeX file without modifying its layout, as if the text of the remarks had been typed over the paper once printed. This is useful to put online copies of papers in your web page which are exactly like the camera-ready ones published in proceedings or journals, but over which some text has been typed, such as a copyright notice, bibliographical information or page numbers.
- MACParrow: a simple LaTeX2e package to add horizontal curved stretchable arrows under and over a formula, as are common in commutative diagrams, like
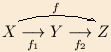 . There are other packages that do this and much more, like xypic, but if you just need over- and underarrows, this one is very compact and simple to use.
. There are other packages that do this and much more, like xypic, but if you just need over- and underarrows, this one is very compact and simple to use.
- www-comand.el package: allows automatic sending of commands to certain WWW sites from inside XEmacs. It currently includes:
and it is straightforward to include others. This package is a minimal modification of Tomasz J. Cholewo's webster-www package that comes in the standard distribution of XEmacs.20.x.
Configuration files for various programs and useful scripts
If you want to use them, be sure to read through them and adapt them to your local configuration; for example, you will need to change some directory names, the email address, check for the existence of certain programs in your system, etc.
- .emacs, for the Emacs or XEmacs editors (requires the www-command.el file)
- init.el and custom.el, internally loaded by my .emacs for the XEmacs editor
- .bibtex, internally loaded by my .emacs for use with XEmacs' BibTeX mode
- .vm, for VM, the XEmacs mail reader
- .ctwmrc, for the CTWM window manager
- llpdf, a shellscript to pass options to ps2pdf so it uses lossless image compression and embeds all fonts, including fonts in Matlab figures
- bitmap-eps, a shellscript to convert an EPS file with lots of objects into a bitmapped EPS file, which will generate smaller PDF files when included in a Latex document
You can get all the files as MACP-config-files.tar.gz.
Miguel A. Carreira-Perpinan
Last modified: Tue Oct 21 23:18:44 PDT 2025
UC Merced |
EECS |
MACP's Home Page
 Java applets
Java applets implementations
implementations LaTeX
LaTeX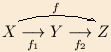 . There are other packages that do this and much more, like xypic, but if you just need over- and underarrows, this one is very compact and simple to use.
. There are other packages that do this and much more, like xypic, but if you just need over- and underarrows, this one is very compact and simple to use. Emacs Lisp
Emacs Lisp